The nmrstarlib Tutorial¶
The nmrstarlib package provides classes and other facilities for parsing,
accessing, and manipulating data stored in NMR-STAR and JSONized NMR-STAR formats.
Also, the nmrstarlib package provides simple command-line interface.
Using nmrstarlib as a library¶
Importing nmrstarlib package¶
If the nmrstarlib package is installed on the system, it can be imported:
In [1]:
import nmrstarlib
Constructing StarFile generator¶
The nmrstarlib module provides the read_files()
generator function that yields StarFile instances. Constructing a
StarFile generator is easy - specify the path to a local NMR-STAR file,
directory of NMR-STAR files, archive of NMR-STAR files or BMRB id:
In [2]:
import nmrstarlib
single_starfile = nmrstarlib.read_files("bmr18569.str") # single NMR-STAR file
starfiles = nmrstarlib.read_files("bmr18569.str", "bmr336.str") # several NMR-STAR files
dir_starfiles = nmrstarlib.read_files("starfiles_dir") # directory of NMR-STAR files
arch_starfiles = nmrstarlib.read_files("starfiles.zip") # archive of NMR-STAR files
url_starfile = nmrstarlib.read_files("18569") # BMRB id of NMR-STAR file
Processing StarFile generator¶
The StarFile generator can be processed in several ways:
- Feed it to a for-loop and process one file at a time:
In [3]:
for starfile in nmrstarlib.read_files("18569", "15000"):
print("BMRB id:", starfile.id) # print BMRB id of StarFile
print("File source:", starfile.source) # print source of StarFile
for saveframe_name in starfile.keys(): # print saveframe names
print("\t", saveframe_name)
BMRB id: 18569
File source: http://rest.bmrb.wisc.edu/bmrb/NMR-STAR3/18569
data
comment_0
save_entry_information
comment_1
save_entry_citation
comment_2
save_assembly
comment_3
save_EVH1
comment_4
save_natural_source
comment_5
save_experimental_source
comment_6
comment_7
save_sample_1
save_sample_2
save_sample_3
save_sample_4
comment_8
save_sample_conditions_1
save_sample_conditions_2
save_sample_conditions_3
save_sample_conditions_4
comment_9
save_AZARA
save_xwinnmr
save_ANSIG
save_CNS
comment_10
comment_11
save_spectrometer_1
save_spectrometer_2
save_NMR_spectrometer_list
comment_12
save_experiment_list
comment_13
comment_14
comment_15
save_chemical_shift_reference_1
comment_16
comment_17
save_assigned_chem_shift_list_1
comment_18
save_combined_NOESY_peak_list
BMRB id: 15000
File source: http://rest.bmrb.wisc.edu/bmrb/NMR-STAR3/15000
data
comment_0
save_entry_information
comment_1
save_citation_1
comment_2
save_assembly
comment_3
save_F5-Phe-cVHP
comment_4
save_natural_source
comment_5
save_experimental_source
comment_6
save_chem_comp_PHF
comment_7
comment_8
save_unlabeled_sample
save_selectively_labeled_sample
comment_9
save_sample_conditions
comment_10
save_NMRPipe
save_PIPP
save_SPARKY
save_CYANA
save_X-PLOR_NIH
comment_11
comment_12
save_spectrometer_1
save_spectrometer_2
save_spectrometer_3
save_spectrometer_4
save_spectrometer_5
save_spectrometer_6
save_NMR_spectrometer_list
comment_13
save_experiment_list
comment_14
comment_15
comment_16
save_chemical_shift_reference_1
comment_17
comment_18
save_assigned_chem_shift_list_1
Note
Once the generator is consumed, it becomes empty and needs to be created again.
In [4]:
sf_generator = nmrstarlib.read_files("18569", "15000")
starfile1 = next(sf_generator)
starfile2 = next(sf_generator)
Note
Once the generator is consumed, it becomes empty and needs to be created again.
In [5]:
starfiles_list = list(nmrstarlib.read_files("18569", "15000"))
Accessing and manipulating data from a single StarFile¶
Since a StarFile is a Python collections.OrderedDict,
data can be accessed and manipulated as with any regular Python dict object
using bracket accessors.
- Accessing data in
StarFile:
In [7]:
starfile = next(nmrstarlib.read_files("15000"))
# list StarFile-level keys, i.e. saveframe names
list(starfile.keys())
Out[7]:
['data',
'comment_0',
'save_entry_information',
'comment_1',
'save_citation_1',
'comment_2',
'save_assembly',
'comment_3',
'save_F5-Phe-cVHP',
'comment_4',
'save_natural_source',
'comment_5',
'save_experimental_source',
'comment_6',
'save_chem_comp_PHF',
'comment_7',
'comment_8',
'save_unlabeled_sample',
'save_selectively_labeled_sample',
'comment_9',
'save_sample_conditions',
'comment_10',
'save_NMRPipe',
'save_PIPP',
'save_SPARKY',
'save_CYANA',
'save_X-PLOR_NIH',
'comment_11',
'comment_12',
'save_spectrometer_1',
'save_spectrometer_2',
'save_spectrometer_3',
'save_spectrometer_4',
'save_spectrometer_5',
'save_spectrometer_6',
'save_NMR_spectrometer_list',
'comment_13',
'save_experiment_list',
'comment_14',
'comment_15',
'comment_16',
'save_chemical_shift_reference_1',
'comment_17',
'comment_18',
'save_assigned_chem_shift_list_1']
In [8]:
# access "data" field
starfile["data"]
Out[8]:
'15000'
In [9]:
# access saveframe
starfile["save_entry_information"]
Out[9]:
OrderedDict([('Entry.Sf_category', 'entry_information'),
('Entry.Sf_framecode', 'entry_information'),
('Entry.ID', '15000'),
('Entry.Title',
'\nSolution structure of chicken villin headpiece subdomain containing a fluorinated side chain in the core\n'),
('Entry.Type', 'macromolecule'),
('Entry.Version_type', 'original'),
('Entry.Submission_date', '2006-09-07'),
('Entry.Accession_date', '2006-09-07'),
('Entry.Last_release_date', '.'),
('Entry.Original_release_date', '.'),
('Entry.Origination', 'author'),
('Entry.NMR_STAR_version', '3.1.1.61'),
('Entry.Original_NMR_STAR_version', '.'),
('Entry.Experimental_method', 'NMR'),
('Entry.Experimental_method_subtype', 'solution'),
('Entry.Details', '.'),
('Entry.BMRB_internal_directory_name', '.'),
('loop_0',
(['Entry_author.Ordinal',
'Entry_author.Given_name',
'Entry_author.Family_name',
'Entry_author.First_initial',
'Entry_author.Middle_initials',
'Entry_author.Family_title',
'Entry_author.Entry_ID'],
[OrderedDict([('Entry_author.Ordinal', '1'),
('Entry_author.Given_name', 'Claudia'),
('Entry_author.Family_name', 'Cornilescu'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'C.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '2'),
('Entry_author.Given_name', 'Gabriel'),
('Entry_author.Family_name', 'Cornilescu'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', '.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '3'),
('Entry_author.Given_name', 'Erik'),
('Entry_author.Family_name', 'Hadley'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'B.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '4'),
('Entry_author.Given_name', 'Samuel'),
('Entry_author.Family_name', 'Gellman'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'H.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '5'),
('Entry_author.Given_name', 'John'),
('Entry_author.Family_name', 'Markley'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'L.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')])])),
('loop_1',
(['SG_project.SG_project_ID',
'SG_project.Project_name',
'SG_project.Full_name_of_center',
'SG_project.Initial_of_center',
'SG_project.Entry_ID'],
[OrderedDict([('SG_project.SG_project_ID', '1'),
('SG_project.Project_name', 'not applicable'),
('SG_project.Full_name_of_center',
'not applicable'),
('SG_project.Initial_of_center', '.'),
('SG_project.Entry_ID', '15000')])])),
('loop_2',
(['Struct_keywords.Keywords',
'Struct_keywords.Text',
'Struct_keywords.Entry_ID'],
[OrderedDict([('Struct_keywords.Keywords',
'chicken villin headpiece'),
('Struct_keywords.Text', '.'),
('Struct_keywords.Entry_ID', '15000')]),
OrderedDict([('Struct_keywords.Keywords', 'fluorinated Phe'),
('Struct_keywords.Text', '.'),
('Struct_keywords.Entry_ID', '15000')]),
OrderedDict([('Struct_keywords.Keywords', 'VHP'),
('Struct_keywords.Text', '.'),
('Struct_keywords.Entry_ID', '15000')])])),
('loop_3',
(['Data_set.Type', 'Data_set.Count', 'Data_set.Entry_ID'],
[OrderedDict([('Data_set.Type', 'assigned_chemical_shifts'),
('Data_set.Count', '1'),
('Data_set.Entry_ID', '15000')])])),
('loop_4',
(['Datum.Type', 'Datum.Count', 'Datum.Entry_ID'],
[OrderedDict([('Datum.Type', '13C chemical shifts'),
('Datum.Count', '77'),
('Datum.Entry_ID', '15000')]),
OrderedDict([('Datum.Type', '15N chemical shifts'),
('Datum.Count', '40'),
('Datum.Entry_ID', '15000')]),
OrderedDict([('Datum.Type', '1H chemical shifts'),
('Datum.Count', '223'),
('Datum.Entry_ID', '15000')])])),
('loop_5',
(['Release.Release_number',
'Release.Format_type',
'Release.Format_version',
'Release.Date',
'Release.Submission_date',
'Release.Type',
'Release.Author',
'Release.Detail',
'Release.Entry_ID'],
[OrderedDict([('Release.Release_number', '2'),
('Release.Format_type', '.'),
('Release.Format_version', '.'),
('Release.Date', '2008-07-17'),
('Release.Submission_date', '2006-09-06'),
('Release.Type', 'update'),
('Release.Author', 'BMRB'),
('Release.Detail', 'complete entry citation'),
('Release.Entry_ID', '15000')]),
OrderedDict([('Release.Release_number', '1'),
('Release.Format_type', '.'),
('Release.Format_version', '.'),
('Release.Date', '2006-10-20'),
('Release.Submission_date', '2006-09-06'),
('Release.Type', 'original'),
('Release.Author', 'author'),
('Release.Detail', 'original release'),
('Release.Entry_ID', '15000')])])),
('loop_6',
(['Related_entries.Database_name',
'Related_entries.Database_accession_code',
'Related_entries.Relationship',
'Related_entries.Entry_ID'],
[OrderedDict([('Related_entries.Database_name', 'PDB'),
('Related_entries.Database_accession_code',
'2JM0'),
('Related_entries.Relationship',
'BMRB Entry Tracking System'),
('Related_entries.Entry_ID', '15000')])]))])
In [10]:
# list saveframe-level keys
list(starfile["save_entry_information"].keys())
Out[10]:
['Entry.Sf_category',
'Entry.Sf_framecode',
'Entry.ID',
'Entry.Title',
'Entry.Type',
'Entry.Version_type',
'Entry.Submission_date',
'Entry.Accession_date',
'Entry.Last_release_date',
'Entry.Original_release_date',
'Entry.Origination',
'Entry.NMR_STAR_version',
'Entry.Original_NMR_STAR_version',
'Entry.Experimental_method',
'Entry.Experimental_method_subtype',
'Entry.Details',
'Entry.BMRB_internal_directory_name',
'loop_0',
'loop_1',
'loop_2',
'loop_3',
'loop_4',
'loop_5',
'loop_6']
In [11]:
# access 'key-value' pairs within saveframes
starfile["save_entry_information"]["Entry.Submission_date"]
Out[11]:
'2006-09-07'
In [12]:
# access loops
starfile["save_entry_information"]["loop_0"]
Out[12]:
(['Entry_author.Ordinal',
'Entry_author.Given_name',
'Entry_author.Family_name',
'Entry_author.First_initial',
'Entry_author.Middle_initials',
'Entry_author.Family_title',
'Entry_author.Entry_ID'],
[OrderedDict([('Entry_author.Ordinal', '1'),
('Entry_author.Given_name', 'Claudia'),
('Entry_author.Family_name', 'Cornilescu'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'C.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '2'),
('Entry_author.Given_name', 'Gabriel'),
('Entry_author.Family_name', 'Cornilescu'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', '.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '3'),
('Entry_author.Given_name', 'Erik'),
('Entry_author.Family_name', 'Hadley'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'B.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '4'),
('Entry_author.Given_name', 'Samuel'),
('Entry_author.Family_name', 'Gellman'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'H.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '5'),
('Entry_author.Given_name', 'John'),
('Entry_author.Family_name', 'Markley'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'L.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')])])
In [13]:
# list loop-level fields
starfile["save_entry_information"]["loop_0"][0]
Out[13]:
['Entry_author.Ordinal',
'Entry_author.Given_name',
'Entry_author.Family_name',
'Entry_author.First_initial',
'Entry_author.Middle_initials',
'Entry_author.Family_title',
'Entry_author.Entry_ID']
In [14]:
# list loop-level values (list of dictionaries)
starfile["save_entry_information"]["loop_0"][1]
Out[14]:
[OrderedDict([('Entry_author.Ordinal', '1'),
('Entry_author.Given_name', 'Claudia'),
('Entry_author.Family_name', 'Cornilescu'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'C.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '2'),
('Entry_author.Given_name', 'Gabriel'),
('Entry_author.Family_name', 'Cornilescu'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', '.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '3'),
('Entry_author.Given_name', 'Erik'),
('Entry_author.Family_name', 'Hadley'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'B.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '4'),
('Entry_author.Given_name', 'Samuel'),
('Entry_author.Family_name', 'Gellman'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'H.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')]),
OrderedDict([('Entry_author.Ordinal', '5'),
('Entry_author.Given_name', 'John'),
('Entry_author.Family_name', 'Markley'),
('Entry_author.First_initial', '.'),
('Entry_author.Middle_initials', 'L.'),
('Entry_author.Family_title', '.'),
('Entry_author.Entry_ID', '15000')])]
In [15]:
# every loop entry is accessed by index
starfile["save_entry_information"]["loop_0"][1][0]["Entry_author.Family_name"]
Out[15]:
'Cornilescu'
- Manipulating data in a
StarFileis easy - access data using bracket accessors and set a new value:
In [16]:
# check submission date
starfile["save_entry_information"]["Entry.Submission_date"]
Out[16]:
'2006-09-07'
In [17]:
# change submission date
starfile["save_entry_information"]["Entry.Submission_date"] = "2015-07-05"
In [18]:
# check that submission date is updated
starfile["save_entry_information"]["Entry.Submission_date"]
Out[18]:
'2015-07-05'
- Printing a
StarFileand its components (saveframe and loop data):
In [19]:
starfile = next(nmrstarlib.read_files("bmr15000.str"))
In [20]:
starfile.print_file(file_format="nmrstar")
data_15000
#######################
# Entry information #
#######################
save_entry_information
_Entry.Sf_category entry_information
_Entry.Sf_framecode entry_information
_Entry.ID 15000
_Entry.Title
;
Solution structure of chicken villin headpiece subdomain containing a fluorinated side chain in the core
;
_Entry.Type macromolecule
_Entry.Version_type original
_Entry.Submission_date 2006-09-07
_Entry.Accession_date 2006-09-07
_Entry.Last_release_date .
_Entry.Original_release_date .
_Entry.Origination author
_Entry.NMR_STAR_version 3.1.1.61
_Entry.Original_NMR_STAR_version .
_Entry.Experimental_method NMR
_Entry.Experimental_method_subtype solution
_Entry.Details .
_Entry.BMRB_internal_directory_name .
loop_
_Entry_author.Ordinal
_Entry_author.Given_name
_Entry_author.Family_name
_Entry_author.First_initial
_Entry_author.Middle_initials
_Entry_author.Family_title
_Entry_author.Entry_ID
1 Claudia Cornilescu . C. . 15000
2 Gabriel Cornilescu . . . 15000
3 Erik Hadley . B. . 15000
4 Samuel Gellman . H. . 15000
5 John Markley . L. . 15000
stop_
loop_
_Datum.Type
_Datum.Count
_Datum.Entry_ID
'13C chemical shifts' 77 15000
'15N chemical shifts' 40 15000
'1H chemical shifts' 223 15000
stop_
save_
save_assigned_chem_shift_list_1
_Assigned_chem_shift_list.Sf_category assigned_chemical_shifts
_Assigned_chem_shift_list.Sf_framecode assigned_chem_shift_list_1
_Assigned_chem_shift_list.Entry_ID 15000
_Assigned_chem_shift_list.ID 1
_Assigned_chem_shift_list.Sample_condition_list_ID 1
_Assigned_chem_shift_list.Sample_condition_list_label $sample_conditions
_Assigned_chem_shift_list.Chem_shift_reference_ID 1
_Assigned_chem_shift_list.Chem_shift_reference_label $chemical_shift_reference_1
_Assigned_chem_shift_list.Chem_shift_1H_err .
_Assigned_chem_shift_list.Chem_shift_13C_err .
_Assigned_chem_shift_list.Chem_shift_15N_err .
_Assigned_chem_shift_list.Chem_shift_31P_err .
_Assigned_chem_shift_list.Chem_shift_2H_err .
_Assigned_chem_shift_list.Chem_shift_19F_err .
_Assigned_chem_shift_list.Error_derivation_method .
_Assigned_chem_shift_list.Details .
_Assigned_chem_shift_list.Text_data_format .
_Assigned_chem_shift_list.Text_data .
loop_
_Atom_chem_shift.ID
_Atom_chem_shift.Assembly_atom_ID
_Atom_chem_shift.Entity_assembly_ID
_Atom_chem_shift.Entity_ID
_Atom_chem_shift.Comp_index_ID
_Atom_chem_shift.Seq_ID
_Atom_chem_shift.Comp_ID
_Atom_chem_shift.Atom_ID
_Atom_chem_shift.Atom_type
_Atom_chem_shift.Atom_isotope_number
_Atom_chem_shift.Val
_Atom_chem_shift.Val_err
_Atom_chem_shift.Assign_fig_of_merit
_Atom_chem_shift.Ambiguity_code
_Atom_chem_shift.Occupancy
_Atom_chem_shift.Resonance_ID
_Atom_chem_shift.Auth_entity_assembly_ID
_Atom_chem_shift.Auth_asym_ID
_Atom_chem_shift.Auth_seq_ID
_Atom_chem_shift.Auth_comp_ID
_Atom_chem_shift.Auth_atom_ID
_Atom_chem_shift.Details
_Atom_chem_shift.Entry_ID
_Atom_chem_shift.Assigned_chem_shift_list_ID
1 . 1 1 2 2 SER H H 1 9.3070 0.01 . . . . . . 2 SER H . 15000 1
2 . 1 1 2 2 SER HA H 1 4.5970 0.01 . . . . . . 2 SER HA . 15000 1
3 . 1 1 2 2 SER HB2 H 1 4.3010 0.01 . . . . . . 2 SER HB2 . 15000 1
4 . 1 1 2 2 SER HB3 H 1 4.0550 0.01 . . . . . . 2 SER HB3 . 15000 1
5 . 1 1 2 2 SER CB C 13 64.6000 0.1 . . . . . . 2 SER CB . 15000 1
6 . 1 1 2 2 SER N N 15 121.5800 0.1 . . . . . . 2 SER N . 15000 1
7 . 1 1 3 3 ASP H H 1 8.0740 0.01 . . . . . . 3 ASP H . 15000 1
8 . 1 1 3 3 ASP HA H 1 4.5580 0.01 . . . . . . 3 ASP HA . 15000 1
9 . 1 1 3 3 ASP HB2 H 1 2.835 0.01 . . . . . . 3 ASP HB2 . 15000 1
10 . 1 1 3 3 ASP HB3 H 1 2.754 0.01 . . . . . . 3 ASP HB3 . 15000 1
11 . 1 1 3 3 ASP CA C 13 57.6400 0.1 . . . . . . 3 ASP CA . 15000 1
12 . 1 1 3 3 ASP N N 15 121.1040 0.1 . . . . . . 3 ASP N . 15000 1
13 . 1 1 4 4 GLU H H 1 8.6520 0.01 . . . . . . 4 GLU H . 15000 1
14 . 1 1 4 4 GLU HA H 1 4.1420 0.01 . . . . . . 4 GLU HA . 15000 1
15 . 1 1 4 4 GLU HB2 H 1 2.0520 0.01 . . . . . . 4 GLU HB2 . 15000 1
16 . 1 1 4 4 GLU HB3 H 1 2.0320 0.01 . . . . . . 4 GLU HB3 . 15000 1
17 . 1 1 4 4 GLU HG2 H 1 2.4540 0.01 . . . . . . 4 GLU HG2 . 15000 1
18 . 1 1 4 4 GLU CB C 13 28.1200 0.1 . . . . . . 4 GLU CB . 15000 1
19 . 1 1 4 4 GLU CG C 13 33.2720 0.1 . . . . . . 4 GLU CG . 15000 1
20 . 1 1 4 4 GLU N N 15 119.8900 0.1 . . . . . . 4 GLU N . 15000 1
stop_
save_
In [21]:
starfile.print_file(file_format="json")
{
"data": "15000",
"comment_0": "#######################\n# Entry information #\n#######################\n",
"save_entry_information": {
"Entry.Sf_category": "entry_information",
"Entry.Sf_framecode": "entry_information",
"Entry.ID": "15000",
"Entry.Title": "\nSolution structure of chicken villin headpiece subdomain containing a fluorinated side chain in the core\n",
"Entry.Type": "macromolecule",
"Entry.Version_type": "original",
"Entry.Submission_date": "2006-09-07",
"Entry.Accession_date": "2006-09-07",
"Entry.Last_release_date": ".",
"Entry.Original_release_date": ".",
"Entry.Origination": "author",
"Entry.NMR_STAR_version": "3.1.1.61",
"Entry.Original_NMR_STAR_version": ".",
"Entry.Experimental_method": "NMR",
"Entry.Experimental_method_subtype": "solution",
"Entry.Details": ".",
"Entry.BMRB_internal_directory_name": ".",
"loop_0": [
[
"Entry_author.Ordinal",
"Entry_author.Given_name",
"Entry_author.Family_name",
"Entry_author.First_initial",
"Entry_author.Middle_initials",
"Entry_author.Family_title",
"Entry_author.Entry_ID"
],
[
{
"Entry_author.Ordinal": "1",
"Entry_author.Given_name": "Claudia",
"Entry_author.Family_name": "Cornilescu",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "C.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "2",
"Entry_author.Given_name": "Gabriel",
"Entry_author.Family_name": "Cornilescu",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": ".",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "3",
"Entry_author.Given_name": "Erik",
"Entry_author.Family_name": "Hadley",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "B.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "4",
"Entry_author.Given_name": "Samuel",
"Entry_author.Family_name": "Gellman",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "H.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "5",
"Entry_author.Given_name": "John",
"Entry_author.Family_name": "Markley",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "L.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
}
]
],
"loop_1": [
[
"Datum.Type",
"Datum.Count",
"Datum.Entry_ID"
],
[
{
"Datum.Type": "13C chemical shifts",
"Datum.Count": "77",
"Datum.Entry_ID": "15000"
},
{
"Datum.Type": "15N chemical shifts",
"Datum.Count": "40",
"Datum.Entry_ID": "15000"
},
{
"Datum.Type": "1H chemical shifts",
"Datum.Count": "223",
"Datum.Entry_ID": "15000"
}
]
]
},
"save_assigned_chem_shift_list_1": {
"Assigned_chem_shift_list.Sf_category": "assigned_chemical_shifts",
"Assigned_chem_shift_list.Sf_framecode": "assigned_chem_shift_list_1",
"Assigned_chem_shift_list.Entry_ID": "15000",
"Assigned_chem_shift_list.ID": "1",
"Assigned_chem_shift_list.Sample_condition_list_ID": "1",
"Assigned_chem_shift_list.Sample_condition_list_label": "$sample_conditions",
"Assigned_chem_shift_list.Chem_shift_reference_ID": "1",
"Assigned_chem_shift_list.Chem_shift_reference_label": "$chemical_shift_reference_1",
"Assigned_chem_shift_list.Chem_shift_1H_err": ".",
"Assigned_chem_shift_list.Chem_shift_13C_err": ".",
"Assigned_chem_shift_list.Chem_shift_15N_err": ".",
"Assigned_chem_shift_list.Chem_shift_31P_err": ".",
"Assigned_chem_shift_list.Chem_shift_2H_err": ".",
"Assigned_chem_shift_list.Chem_shift_19F_err": ".",
"Assigned_chem_shift_list.Error_derivation_method": ".",
"Assigned_chem_shift_list.Details": ".",
"Assigned_chem_shift_list.Text_data_format": ".",
"Assigned_chem_shift_list.Text_data": ".",
"loop_0": [
[
"Atom_chem_shift.ID",
"Atom_chem_shift.Assembly_atom_ID",
"Atom_chem_shift.Entity_assembly_ID",
"Atom_chem_shift.Entity_ID",
"Atom_chem_shift.Comp_index_ID",
"Atom_chem_shift.Seq_ID",
"Atom_chem_shift.Comp_ID",
"Atom_chem_shift.Atom_ID",
"Atom_chem_shift.Atom_type",
"Atom_chem_shift.Atom_isotope_number",
"Atom_chem_shift.Val",
"Atom_chem_shift.Val_err",
"Atom_chem_shift.Assign_fig_of_merit",
"Atom_chem_shift.Ambiguity_code",
"Atom_chem_shift.Occupancy",
"Atom_chem_shift.Resonance_ID",
"Atom_chem_shift.Auth_entity_assembly_ID",
"Atom_chem_shift.Auth_asym_ID",
"Atom_chem_shift.Auth_seq_ID",
"Atom_chem_shift.Auth_comp_ID",
"Atom_chem_shift.Auth_atom_ID",
"Atom_chem_shift.Details",
"Atom_chem_shift.Entry_ID",
"Atom_chem_shift.Assigned_chem_shift_list_ID"
],
[
{
"Atom_chem_shift.ID": "1",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "2",
"Atom_chem_shift.Seq_ID": "2",
"Atom_chem_shift.Comp_ID": "SER",
"Atom_chem_shift.Atom_ID": "H",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "9.3070",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "2",
"Atom_chem_shift.Auth_comp_ID": "SER",
"Atom_chem_shift.Auth_atom_ID": "H",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "2",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "2",
"Atom_chem_shift.Seq_ID": "2",
"Atom_chem_shift.Comp_ID": "SER",
"Atom_chem_shift.Atom_ID": "HA",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "4.5970",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "2",
"Atom_chem_shift.Auth_comp_ID": "SER",
"Atom_chem_shift.Auth_atom_ID": "HA",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "3",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "2",
"Atom_chem_shift.Seq_ID": "2",
"Atom_chem_shift.Comp_ID": "SER",
"Atom_chem_shift.Atom_ID": "HB2",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "4.3010",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "2",
"Atom_chem_shift.Auth_comp_ID": "SER",
"Atom_chem_shift.Auth_atom_ID": "HB2",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "4",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "2",
"Atom_chem_shift.Seq_ID": "2",
"Atom_chem_shift.Comp_ID": "SER",
"Atom_chem_shift.Atom_ID": "HB3",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "4.0550",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "2",
"Atom_chem_shift.Auth_comp_ID": "SER",
"Atom_chem_shift.Auth_atom_ID": "HB3",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "5",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "2",
"Atom_chem_shift.Seq_ID": "2",
"Atom_chem_shift.Comp_ID": "SER",
"Atom_chem_shift.Atom_ID": "CB",
"Atom_chem_shift.Atom_type": "C",
"Atom_chem_shift.Atom_isotope_number": "13",
"Atom_chem_shift.Val": "64.6000",
"Atom_chem_shift.Val_err": "0.1",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "2",
"Atom_chem_shift.Auth_comp_ID": "SER",
"Atom_chem_shift.Auth_atom_ID": "CB",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "6",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "2",
"Atom_chem_shift.Seq_ID": "2",
"Atom_chem_shift.Comp_ID": "SER",
"Atom_chem_shift.Atom_ID": "N",
"Atom_chem_shift.Atom_type": "N",
"Atom_chem_shift.Atom_isotope_number": "15",
"Atom_chem_shift.Val": "121.5800",
"Atom_chem_shift.Val_err": "0.1",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "2",
"Atom_chem_shift.Auth_comp_ID": "SER",
"Atom_chem_shift.Auth_atom_ID": "N",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "7",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "3",
"Atom_chem_shift.Seq_ID": "3",
"Atom_chem_shift.Comp_ID": "ASP",
"Atom_chem_shift.Atom_ID": "H",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "8.0740",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "3",
"Atom_chem_shift.Auth_comp_ID": "ASP",
"Atom_chem_shift.Auth_atom_ID": "H",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "8",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "3",
"Atom_chem_shift.Seq_ID": "3",
"Atom_chem_shift.Comp_ID": "ASP",
"Atom_chem_shift.Atom_ID": "HA",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "4.5580",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "3",
"Atom_chem_shift.Auth_comp_ID": "ASP",
"Atom_chem_shift.Auth_atom_ID": "HA",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "9",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "3",
"Atom_chem_shift.Seq_ID": "3",
"Atom_chem_shift.Comp_ID": "ASP",
"Atom_chem_shift.Atom_ID": "HB2",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "2.835",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "3",
"Atom_chem_shift.Auth_comp_ID": "ASP",
"Atom_chem_shift.Auth_atom_ID": "HB2",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "10",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "3",
"Atom_chem_shift.Seq_ID": "3",
"Atom_chem_shift.Comp_ID": "ASP",
"Atom_chem_shift.Atom_ID": "HB3",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "2.754",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "3",
"Atom_chem_shift.Auth_comp_ID": "ASP",
"Atom_chem_shift.Auth_atom_ID": "HB3",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "11",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "3",
"Atom_chem_shift.Seq_ID": "3",
"Atom_chem_shift.Comp_ID": "ASP",
"Atom_chem_shift.Atom_ID": "CA",
"Atom_chem_shift.Atom_type": "C",
"Atom_chem_shift.Atom_isotope_number": "13",
"Atom_chem_shift.Val": "57.6400",
"Atom_chem_shift.Val_err": "0.1",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "3",
"Atom_chem_shift.Auth_comp_ID": "ASP",
"Atom_chem_shift.Auth_atom_ID": "CA",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "12",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "3",
"Atom_chem_shift.Seq_ID": "3",
"Atom_chem_shift.Comp_ID": "ASP",
"Atom_chem_shift.Atom_ID": "N",
"Atom_chem_shift.Atom_type": "N",
"Atom_chem_shift.Atom_isotope_number": "15",
"Atom_chem_shift.Val": "121.1040",
"Atom_chem_shift.Val_err": "0.1",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "3",
"Atom_chem_shift.Auth_comp_ID": "ASP",
"Atom_chem_shift.Auth_atom_ID": "N",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "13",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "4",
"Atom_chem_shift.Seq_ID": "4",
"Atom_chem_shift.Comp_ID": "GLU",
"Atom_chem_shift.Atom_ID": "H",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "8.6520",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "4",
"Atom_chem_shift.Auth_comp_ID": "GLU",
"Atom_chem_shift.Auth_atom_ID": "H",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "14",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "4",
"Atom_chem_shift.Seq_ID": "4",
"Atom_chem_shift.Comp_ID": "GLU",
"Atom_chem_shift.Atom_ID": "HA",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "4.1420",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "4",
"Atom_chem_shift.Auth_comp_ID": "GLU",
"Atom_chem_shift.Auth_atom_ID": "HA",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "15",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "4",
"Atom_chem_shift.Seq_ID": "4",
"Atom_chem_shift.Comp_ID": "GLU",
"Atom_chem_shift.Atom_ID": "HB2",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "2.0520",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "4",
"Atom_chem_shift.Auth_comp_ID": "GLU",
"Atom_chem_shift.Auth_atom_ID": "HB2",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "16",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "4",
"Atom_chem_shift.Seq_ID": "4",
"Atom_chem_shift.Comp_ID": "GLU",
"Atom_chem_shift.Atom_ID": "HB3",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "2.0320",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "4",
"Atom_chem_shift.Auth_comp_ID": "GLU",
"Atom_chem_shift.Auth_atom_ID": "HB3",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "17",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "4",
"Atom_chem_shift.Seq_ID": "4",
"Atom_chem_shift.Comp_ID": "GLU",
"Atom_chem_shift.Atom_ID": "HG2",
"Atom_chem_shift.Atom_type": "H",
"Atom_chem_shift.Atom_isotope_number": "1",
"Atom_chem_shift.Val": "2.4540",
"Atom_chem_shift.Val_err": "0.01",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "4",
"Atom_chem_shift.Auth_comp_ID": "GLU",
"Atom_chem_shift.Auth_atom_ID": "HG2",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "18",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "4",
"Atom_chem_shift.Seq_ID": "4",
"Atom_chem_shift.Comp_ID": "GLU",
"Atom_chem_shift.Atom_ID": "CB",
"Atom_chem_shift.Atom_type": "C",
"Atom_chem_shift.Atom_isotope_number": "13",
"Atom_chem_shift.Val": "28.1200",
"Atom_chem_shift.Val_err": "0.1",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "4",
"Atom_chem_shift.Auth_comp_ID": "GLU",
"Atom_chem_shift.Auth_atom_ID": "CB",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "19",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "4",
"Atom_chem_shift.Seq_ID": "4",
"Atom_chem_shift.Comp_ID": "GLU",
"Atom_chem_shift.Atom_ID": "CG",
"Atom_chem_shift.Atom_type": "C",
"Atom_chem_shift.Atom_isotope_number": "13",
"Atom_chem_shift.Val": "33.2720",
"Atom_chem_shift.Val_err": "0.1",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "4",
"Atom_chem_shift.Auth_comp_ID": "GLU",
"Atom_chem_shift.Auth_atom_ID": "CG",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
},
{
"Atom_chem_shift.ID": "20",
"Atom_chem_shift.Assembly_atom_ID": ".",
"Atom_chem_shift.Entity_assembly_ID": "1",
"Atom_chem_shift.Entity_ID": "1",
"Atom_chem_shift.Comp_index_ID": "4",
"Atom_chem_shift.Seq_ID": "4",
"Atom_chem_shift.Comp_ID": "GLU",
"Atom_chem_shift.Atom_ID": "N",
"Atom_chem_shift.Atom_type": "N",
"Atom_chem_shift.Atom_isotope_number": "15",
"Atom_chem_shift.Val": "119.8900",
"Atom_chem_shift.Val_err": "0.1",
"Atom_chem_shift.Assign_fig_of_merit": ".",
"Atom_chem_shift.Ambiguity_code": ".",
"Atom_chem_shift.Occupancy": ".",
"Atom_chem_shift.Resonance_ID": ".",
"Atom_chem_shift.Auth_entity_assembly_ID": ".",
"Atom_chem_shift.Auth_asym_ID": ".",
"Atom_chem_shift.Auth_seq_ID": "4",
"Atom_chem_shift.Auth_comp_ID": "GLU",
"Atom_chem_shift.Auth_atom_ID": "N",
"Atom_chem_shift.Details": ".",
"Atom_chem_shift.Entry_ID": "15000",
"Atom_chem_shift.Assigned_chem_shift_list_ID": "1"
}
]
]
}
}
In [22]:
starfile.print_saveframe("save_entry_information", file_format="nmrstar")
_Entry.Sf_category entry_information
_Entry.Sf_framecode entry_information
_Entry.ID 15000
_Entry.Title
;
Solution structure of chicken villin headpiece subdomain containing a fluorinated side chain in the core
;
_Entry.Type macromolecule
_Entry.Version_type original
_Entry.Submission_date 2006-09-07
_Entry.Accession_date 2006-09-07
_Entry.Last_release_date .
_Entry.Original_release_date .
_Entry.Origination author
_Entry.NMR_STAR_version 3.1.1.61
_Entry.Original_NMR_STAR_version .
_Entry.Experimental_method NMR
_Entry.Experimental_method_subtype solution
_Entry.Details .
_Entry.BMRB_internal_directory_name .
loop_
_Entry_author.Ordinal
_Entry_author.Given_name
_Entry_author.Family_name
_Entry_author.First_initial
_Entry_author.Middle_initials
_Entry_author.Family_title
_Entry_author.Entry_ID
1 Claudia Cornilescu . C. . 15000
2 Gabriel Cornilescu . . . 15000
3 Erik Hadley . B. . 15000
4 Samuel Gellman . H. . 15000
5 John Markley . L. . 15000
stop_
loop_
_Datum.Type
_Datum.Count
_Datum.Entry_ID
'13C chemical shifts' 77 15000
'15N chemical shifts' 40 15000
'1H chemical shifts' 223 15000
stop_
In [23]:
starfile.print_saveframe("save_entry_information", file_format="json")
{
"Entry.Sf_category": "entry_information",
"Entry.Sf_framecode": "entry_information",
"Entry.ID": "15000",
"Entry.Title": "\nSolution structure of chicken villin headpiece subdomain containing a fluorinated side chain in the core\n",
"Entry.Type": "macromolecule",
"Entry.Version_type": "original",
"Entry.Submission_date": "2006-09-07",
"Entry.Accession_date": "2006-09-07",
"Entry.Last_release_date": ".",
"Entry.Original_release_date": ".",
"Entry.Origination": "author",
"Entry.NMR_STAR_version": "3.1.1.61",
"Entry.Original_NMR_STAR_version": ".",
"Entry.Experimental_method": "NMR",
"Entry.Experimental_method_subtype": "solution",
"Entry.Details": ".",
"Entry.BMRB_internal_directory_name": ".",
"loop_0": [
[
"Entry_author.Ordinal",
"Entry_author.Given_name",
"Entry_author.Family_name",
"Entry_author.First_initial",
"Entry_author.Middle_initials",
"Entry_author.Family_title",
"Entry_author.Entry_ID"
],
[
{
"Entry_author.Ordinal": "1",
"Entry_author.Given_name": "Claudia",
"Entry_author.Family_name": "Cornilescu",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "C.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "2",
"Entry_author.Given_name": "Gabriel",
"Entry_author.Family_name": "Cornilescu",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": ".",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "3",
"Entry_author.Given_name": "Erik",
"Entry_author.Family_name": "Hadley",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "B.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "4",
"Entry_author.Given_name": "Samuel",
"Entry_author.Family_name": "Gellman",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "H.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "5",
"Entry_author.Given_name": "John",
"Entry_author.Family_name": "Markley",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "L.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
}
]
],
"loop_1": [
[
"Datum.Type",
"Datum.Count",
"Datum.Entry_ID"
],
[
{
"Datum.Type": "13C chemical shifts",
"Datum.Count": "77",
"Datum.Entry_ID": "15000"
},
{
"Datum.Type": "15N chemical shifts",
"Datum.Count": "40",
"Datum.Entry_ID": "15000"
},
{
"Datum.Type": "1H chemical shifts",
"Datum.Count": "223",
"Datum.Entry_ID": "15000"
}
]
]
}
In [24]:
starfile.print_loop("save_entry_information", "loop_0", file_format="nmrstar")
_Entry_author.Ordinal
_Entry_author.Given_name
_Entry_author.Family_name
_Entry_author.First_initial
_Entry_author.Middle_initials
_Entry_author.Family_title
_Entry_author.Entry_ID
1 Claudia Cornilescu . C. . 15000
2 Gabriel Cornilescu . . . 15000
3 Erik Hadley . B. . 15000
4 Samuel Gellman . H. . 15000
5 John Markley . L. . 15000
In [25]:
starfile.print_loop("save_entry_information", "loop_0", file_format="json")
[
[
"Entry_author.Ordinal",
"Entry_author.Given_name",
"Entry_author.Family_name",
"Entry_author.First_initial",
"Entry_author.Middle_initials",
"Entry_author.Family_title",
"Entry_author.Entry_ID"
],
[
{
"Entry_author.Ordinal": "1",
"Entry_author.Given_name": "Claudia",
"Entry_author.Family_name": "Cornilescu",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "C.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "2",
"Entry_author.Given_name": "Gabriel",
"Entry_author.Family_name": "Cornilescu",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": ".",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "3",
"Entry_author.Given_name": "Erik",
"Entry_author.Family_name": "Hadley",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "B.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "4",
"Entry_author.Given_name": "Samuel",
"Entry_author.Family_name": "Gellman",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "H.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
},
{
"Entry_author.Ordinal": "5",
"Entry_author.Given_name": "John",
"Entry_author.Family_name": "Markley",
"Entry_author.First_initial": ".",
"Entry_author.Middle_initials": "L.",
"Entry_author.Family_title": ".",
"Entry_author.Entry_ID": "15000"
}
]
]
Accessing chemical shift data:
Chemical shift data can be accessed using bracket accessors as described above using a saveframe name and loop name:
In [26]:
starfile["save_assigned_chem_shift_list_1"]["loop_0"][0]
Out[26]:
['Atom_chem_shift.ID',
'Atom_chem_shift.Assembly_atom_ID',
'Atom_chem_shift.Entity_assembly_ID',
'Atom_chem_shift.Entity_ID',
'Atom_chem_shift.Comp_index_ID',
'Atom_chem_shift.Seq_ID',
'Atom_chem_shift.Comp_ID',
'Atom_chem_shift.Atom_ID',
'Atom_chem_shift.Atom_type',
'Atom_chem_shift.Atom_isotope_number',
'Atom_chem_shift.Val',
'Atom_chem_shift.Val_err',
'Atom_chem_shift.Assign_fig_of_merit',
'Atom_chem_shift.Ambiguity_code',
'Atom_chem_shift.Occupancy',
'Atom_chem_shift.Resonance_ID',
'Atom_chem_shift.Auth_entity_assembly_ID',
'Atom_chem_shift.Auth_asym_ID',
'Atom_chem_shift.Auth_seq_ID',
'Atom_chem_shift.Auth_comp_ID',
'Atom_chem_shift.Auth_atom_ID',
'Atom_chem_shift.Details',
'Atom_chem_shift.Entry_ID',
'Atom_chem_shift.Assigned_chem_shift_list_ID']
In [27]:
starfile["save_assigned_chem_shift_list_1"]["loop_0"][1][0]["Atom_chem_shift.Seq_ID"]
Out[27]:
'2'
In [28]:
starfile["save_assigned_chem_shift_list_1"]["loop_0"][1][0]["Atom_chem_shift.Comp_ID"]
Out[28]:
'SER'
In [29]:
starfile["save_assigned_chem_shift_list_1"]["loop_0"][1][0]["Atom_chem_shift.Atom_ID"]
Out[29]:
'H'
In [30]:
starfile["save_assigned_chem_shift_list_1"]["loop_0"][1][0]["Atom_chem_shift.Val"]
Out[30]:
'9.3070'
In [31]:
starfile["save_assigned_chem_shift_list_1"]["loop_0"][1][1]["Atom_chem_shift.Atom_ID"]
Out[31]:
'HA'
In [32]:
starfile["save_assigned_chem_shift_list_1"]["loop_0"][1][1]["Atom_chem_shift.Val"]
Out[32]:
'4.5970'
In [33]:
starfile["save_assigned_chem_shift_list_1"]["loop_0"][1][2]["Atom_chem_shift.Atom_ID"]
Out[33]:
'HB2'
In [34]:
starfile["save_assigned_chem_shift_list_1"]["loop_0"][1][2]["Atom_chem_shift.Val"]
Out[34]:
'4.3010'
Also the StarFile class provides a
chem_shifts_by_residue() method that organizes
chemical shits into a list of collections.OrderedDict data structures
(keys - sequence id, values - chemical shift data) - one for each protein chain,
if multiple chains are present within the file:
In [35]:
# access all chemical shifts
starfile.chem_shifts_by_residue()
Out[35]:
[OrderedDict([('2',
OrderedDict([('AA3Code', 'SER'),
('Seq_ID', '2'),
('H', '9.3070'),
('HA', '4.5970'),
('HB2', '4.3010'),
('HB3', '4.0550'),
('CB', '64.6000'),
('N', '121.5800')])),
('3',
OrderedDict([('AA3Code', 'ASP'),
('Seq_ID', '3'),
('H', '8.0740'),
('HA', '4.5580'),
('HB2', '2.835'),
('HB3', '2.754'),
('CA', '57.6400'),
('N', '121.1040')])),
('4',
OrderedDict([('AA3Code', 'GLU'),
('Seq_ID', '4'),
('H', '8.6520'),
('HA', '4.1420'),
('HB2', '2.0520'),
('HB3', '2.0320'),
('HG2', '2.4540'),
('CB', '28.1200'),
('CG', '33.2720'),
('N', '119.8900')]))])]
In [36]:
# access chemical shifts for "SER" and "GLU" amino acids
starfile.chem_shifts_by_residue(amino_acids=["SER", "GLU"])
Out[36]:
[OrderedDict([('2',
OrderedDict([('AA3Code', 'SER'),
('Seq_ID', '2'),
('H', '9.3070'),
('HA', '4.5970'),
('HB2', '4.3010'),
('HB3', '4.0550'),
('CB', '64.6000'),
('N', '121.5800')])),
('4',
OrderedDict([('AA3Code', 'GLU'),
('Seq_ID', '4'),
('H', '8.6520'),
('HA', '4.1420'),
('HB2', '2.0520'),
('HB3', '2.0320'),
('HG2', '2.4540'),
('CB', '28.1200'),
('CG', '33.2720'),
('N', '119.8900')]))])]
In [37]:
# access chemical shifts for "SER" and "GLU" amino acids for "CB" and "CG" atoms
starfile.chem_shifts_by_residue(amino_acids=["SER", "GLU"], atoms=["CB", "CG"])
Out[37]:
[OrderedDict([('2',
OrderedDict([('AA3Code', 'SER'),
('Seq_ID', '2'),
('CB', '64.6000')])),
('4',
OrderedDict([('AA3Code', 'GLU'),
('Seq_ID', '4'),
('CB', '28.1200'),
('CG', '33.2720')]))])]
In [38]:
# acceess chemical shifts for specific amino acid and specific atom
starfile.chem_shifts_by_residue(amino_acids_and_atoms={"SER":["HA", "HB2", "HB3"], "ASP": ["CA", "N"]})
Out[38]:
[OrderedDict([('2',
OrderedDict([('AA3Code', 'SER'),
('Seq_ID', '2'),
('HA', '4.5970'),
('HB2', '4.3010'),
('HB3', '4.0550')])),
('3',
OrderedDict([('AA3Code', 'ASP'),
('Seq_ID', '3'),
('CA', '57.6400'),
('N', '121.1040')]))])]
Writing data from a StarFile object into a file¶
Data from a StarFile can be written into file
in original NMR-STAR format or in equivalent JSON format using
write():
- Writing into a NMR-STAR formatted file:
In [39]:
with open("out/bmr15000_modified.str", "w") as outfile:
starfile.write(outfile, file_format="nmrstar")
- Writing into a JSONized NMR-STAR formatted file:
In [40]:
with open("out/bmr15000_modified.json", "w") as outfile:
starfile.write(outfile, file_format="json")
Converting NMR-STAR files¶
NMR-STAR files can be converted between the NMR-STAR file format and a JSONized NMR-STAR
file format using nmrstarlib.converter and nmrstarlib.translator modules.
One-to-one file conversions¶
- Converting from the NMR-STAR file format into its equivalent JSON file format:
In [41]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToStarFile
# Using valid BMRB id to access file from URL: from_path="18569"
converter = Converter(StarFileToStarFile(from_path="18569", to_path="out/bmr18569.json",
from_format="nmrstar", to_format="json"))
converter.convert()
- Converting from JSON file format into its equivalent NMR-STAR file format:
In [42]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToStarFile
# Using generated above "bmr18569.json" file
converter = Converter(StarFileToStarFile(from_path="bmr18569.json", to_path="out/bmr18569.str",
from_format="json", to_format="nmrstar"))
converter.convert()
Many-to-many files conversions¶
- Converting from the directory of NMR-STAR formatted files into its equivalent JSON formatted files:
In [43]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToStarFile
converter = Converter(StarFileToStarFile(from_path="starfiles_dir_nmrstar", to_path="out/starfiles_dir_json",
from_format="nmrstar", to_format="json"))
converter.convert()
- Converting from the directory of JSONized NMR-STAR formatted files into NMR-STAR formatted files:
In [44]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToStarFile
converter = Converter(StarFileToStarFile(from_path="starfiles_dir_json", to_path="out/starfiles_dir_nmrstar",
from_format="json", to_format="nmrstar"))
converter.convert()
Note
Many-to-many files and one-to-one file conversions are available.
See nmrstarlib.converter for full list of available conversions.
Creating simulated peak lists from NMR-STAR formatted files¶
Creating simulated peak lists without variance¶
Chemical shift values and assignment information deposited in NMR-STAR formatted files can be used to generate a large number of simulated peak lists for different types of solution and solid-state NMR experiments. Many different types of standard NMR experiments are defined in the spectrum_description.json configuration file. We will be using HNcoCACB spectrum type for the following examples.
- Creating a zero-variance HNcoCACB peak list file in sparky-like format from NMR-STAR formatted file:
In [45]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToPeakList
# Using valid BMRB id to access file from URL: from_path="18569"
converter = Converter(StarFileToPeakList(from_path="18569", to_path="out/18569_HNcoCACB.txt",
from_format="nmrstar", to_format="sparky",
spectrum_name="HNcoCACB"))
converter.convert()
The generated 18569_HNcoCACB.txt peak list file should look like the following:
Assignment w1 w2 w3
SER2H-SER2N-MET1CA 8.225 117.197 55.489
SER2H-SER2N-MET1CB 8.225 117.197 32.848
GLU3H-GLU3N-SER2CA 8.002 119.833 58.593
GLU3H-GLU3N-SER2CB 8.002 119.833 64.057
THR4H-THR4N-GLU3CA 8.956 117.212 55.651
THR4H-THR4N-GLU3CB 8.956 117.212 32.952
...
- Creating a zero-variance HNcoCACB peak list file in json format from a NMR-STAR formatted file:
In [46]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToPeakList
# Using valid BMRB id to access file from URL: from_path="18569"
converter = Converter(StarFileToPeakList(from_path="18569", to_path="out/18569_HNcoCACB.json",
from_format="nmrstar", to_format="json",
spectrum_name="HNcoCACB"))
converter.convert()
The generated 18569_HNcoCACB.json peak list file should look like the following:
[
{"Assignment": ["SER2H", "SER2N", "MET1CA"], "Dimensions": [8.225, 117.197, 55.489]},
{"Assignment": ["SER2H", "SER2N", "MET1CB"], "Dimensions": [8.225, 117.197, 32.848]},
{"Assignment": ["GLU3H", "GLU3N", "SER2CA"], "Dimensions": [8.002, 119.833, 58.593]},
{"Assignment": ["GLU3H", "GLU3N", "SER2CB"], "Dimensions": [8.002, 119.833, 64.057]},
{"Assignment": ["THR4H", "THR4N", "GLU3CA"], "Dimensions": [8.956, 117.212, 55.651]},
{"Assignment": ["THR4H", "THR4N", "GLU3CB"], "Dimensions": [8.956, 117.212, 32.952]},
...
]
Creating simulated peak lists with variance drawn from random normal distribution¶
- Creating a HNcoCACB peak list file in sparky-like format and adding noise values to peak dimensions from a single source of variance, i.e. 100% of peaks will have chemical shift values adjusted using noise values from the defined random normal distribution:
In [47]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToPeakList
from nmrstarlib.noise import NoiseGenerator
# create parameters dictionary for random normal distribution
parameters = {"H_loc": [0], "C_loc": [0], "N_loc": [0],
"H_scale": [0.001], "C_scale": [0.01], "N_scale": [0.01]}
# create random normal noise generator
random_normal_noise_generator = NoiseGenerator(parameters)
# Using valid BMRB id to access file from URL: from_path="18569"
converter = Converter(StarFileToPeakList(from_path="18569", to_path="out/18569_HNcoCACB_ssv_HCN.txt",
from_format="nmrstar", to_format="sparky",
spectrum_name="HNcoCACB",
noise_generator=random_normal_noise_generator))
converter.convert()
The generated 18569_HNcoCACB_ssv_HCN.txt peak list file should look like the following:
Assignment w1 w2 w3
SER2H-SER2N-MET1CA 8.226026 117.193655 55.477204
SER2H-SER2N-MET1CB 8.224649 117.184255 32.845212
GLU3H-GLU3N-SER2CA 8.003282 119.841221 58.603253
GLU3H-GLU3N-SER2CB 8.002372 119.827019 64.067278
THR4H-THR4N-GLU3CA 8.955568 117.215237 55.663902
THR4H-THR4N-GLU3CB 8.955757 117.206167 32.96412
- Creating a HNcoCACB peak list file in sparky-like format and adding noise values to H and N peak dimensions but not C peak dimension from a single source of variance, i.e. 100% of peaks will have chemical shift values adjusted using noise values from the defined random normal distribution:
In [48]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToPeakList
from nmrstarlib.noise import NoiseGenerator
# create parameters dictionary for random normal distribution
parameters = {"H_loc": [0], "C_loc": [None], "N_loc": [0],
"H_scale": [0.001], "C_scale": [None], "N_scale": [0.01]}
# create random normal noise generator
random_normal_noise_generator = NoiseGenerator(parameters)
# Using valid BMRB id to access file from URL: from_path="18569"
converter = Converter(StarFileToPeakList(from_path="18569", to_path="out/18569_HNcoCACB_ssv_HN.txt",
from_format="nmrstar", to_format="sparky",
spectrum_name="HNcoCACB",
noise_generator=random_normal_noise_generator))
converter.convert()
The generated 18569_HNcoCACB_ssv_HN.txt peak list file should look like the following (note the chemical shift values differences in H and N dimensions for peaks that belong to the same spin system):
Assignment w1 w2 w3
SER2H-SER2N-MET1CA 8.226085 117.191527 55.489
SER2H-SER2N-MET1CB 8.224509 117.204666 32.848
GLU3H-GLU3N-SER2CA 8.001657 119.846806 58.593
GLU3H-GLU3N-SER2CB 8.003165 119.8268 64.057
THR4H-THR4N-GLU3CA 8.956946 117.209486 55.651
THR4H-THR4N-GLU3CB 8.955755 117.209889 32.952
- Creating a HNcoCACB peak list file in sparky-like format and adding noise values to peak dimensions from two sources of variance, i.e. chemical shift values will be adjusted using noise values from two random normal distributions. In order to specify two sources of variance, we need to provide how we want to split our peak list and provide statistical distribution parameters for both distributions. Let’s say we want 70 % of peaks to have a smaller variance in H and N dimensions and 30 % of peaks to have a larger variance in H and N dimensions:
In [49]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToPeakList
from nmrstarlib.noise import NoiseGenerator
# create parameters dictionary for random normal distribution
parameters = {"H_loc": [0, 0], "C_loc": [None, None], "N_loc": [0, 0],
"H_scale": [0.001, 0.005], "C_scale": [None, None], "N_scale": [0.01, 0.05]}
# create random normal noise generator
random_normal_noise_generator = NoiseGenerator(parameters)
# Using valid BMRB id to access file from URL: from_path="18569"
converter = Converter(StarFileToPeakList(from_path="18569", to_path="out/18569_HNcoCACB_tsv_HN.txt",
from_format="nmrstar", to_format="sparky",
spectrum_name="HNcoCACB",
plsplit=(70,30),
noise_generator=random_normal_noise_generator))
converter.convert()
The generated 18569.txt peak list file should look like the following (note the larger variance in the last four peaks especially in N dimension):
Assignment w1 w2 w3
SER2H-SER2N-MET1CA 8.223356 117.208041 55.489
SER2H-SER2N-MET1CB 8.22532 117.184278 32.848
GLU3H-GLU3N-SER2CA 8.00271 119.847153 58.593
GLU3H-GLU3N-SER2CB 8.002822 119.824752 64.057
...
GLU114H-GLU114N-LEU113CA 7.614195 118.672897 56.14
GLU114H-GLU114N-LEU113CB 7.628722 118.565859 43.249
GLY115H-GLY115N-GLU114CA 7.583248 113.45153 57.005
GLY115H-GLY115N-GLU114CB 7.596634 113.472049 30.079
Creating simulated peak lists with variance drawn from other distribution types¶
- It is also possible to generate the simulated peak lists using other
types of statistical distribution functions. For example, let’s
simulate the peak list using noise values drawn from
chisquaredistribution for 5 degrees of freedom for H and N dimensions from single source of variance.
In [50]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToPeakList
from nmrstarlib.noise import NoiseGenerator
# create parameters dictionary for distribution
parameters = {"H_df": [5], "C_df": [None], "N_df": [5]}
# create chisquare noise generator
chisquare_noise_generator = NoiseGenerator(parameters, distribution_name="chisquare")
# Using valid BMRB id to access file from URL: from_path="18569"
converter = Converter(StarFileToPeakList(from_path="18569", to_path="out/18569_HNcoCACB_ssv_HN_chi2.txt",
from_format="nmrstar", to_format="sparky",
spectrum_name="HNcoCACB",
noise_generator=chisquare_noise_generator))
converter.convert()
The generated 18569_HNcoCACB_ssv_HN_chi2.txt peak list file should look like the following:
Assignment w1 w2 w3
SER2H-SER2N-MET1CA 12.50083 127.197738 55.489
SER2H-SER2N-MET1CB 10.495158 121.039655 32.848
GLU3H-GLU3N-SER2CA 15.597162 124.603078 58.593
GLU3H-GLU3N-SER2CB 8.340404 126.784481 64.057
THR4H-THR4N-GLU3CA 10.010804 120.476893 55.651
THR4H-THR4N-GLU3CB 11.961498 121.681636 32.952
- Below is the list of all supported distribution functions along with their parameters
if the
numpylibrary is not installed:
{
{"function": "uniform", "parameters": ["low", "high"]},
{"function": "triangular", "parameters": ["left", "right", "mode"]},
{"function": "beta", "parameters": ["a", "b"]},
{"function": "exponential", "parameters": ["scale"]},
{"function": "gamma", "parameters": ["shape", "scale"]},
{"function": "gauss", "parameters": ["mu", "sigma"]},
{"function": "normal", "parameters": ["loc", "scale"]},
{"function": "lognormal", "parameters": ["mean", "sigma"]},
{"function": "vonmises", "parameters": ["mu", "kappa"]},
{"function": "pareto", "parameters": ["a"]}
}
- And the list of all supported distribution functions along with their parameters
if the
numpylibrary is installed:
{
{"function": "beta", "parameters": ["a", "b"]},
{"function": "binomial", "parameters": ["n", "p"]},
{"function": "chisquare", "parameters": ["df"]},
{"function": "exponential", "parameters": ["scale"]},
{"function": "f", "parameters": ["dfnum", "dfden"]},
{"function": "gamma", "parameters": ["shape", "scale"]},
{"function": "geometric", "parameters": ["p"]},
{"function": "gumbel", "parameters": ["loc", "scale"]},
{"function": "hypergeometric", "parameters": ["ngood", "nbad", "nsample"]},
{"function": "laplace", "parameters": ["loc", "scale"]},
{"function": "logistic", "parameters": ["loc", "scale"]},
{"function": "lognormal", "parameters": ["mean", "sigma"]},
{"function": "logseries", "parameters": ["p"]},
{"function": "negative_binomial", "parameters": ["n", "p"]},
{"function": "noncentral_chisquare", "parameters": ["df", "nonc"]},
{"function": "noncentral_f", "parameters": ["dfnum", "dfden", "nonc"]},
{"function": "normal", "parameters": ["loc", "scale"]},
{"function": "pareto", "parameters": ["a"]},
{"function": "poisson", "parameters": ["lam"]},
{"function": "power", "parameters": ["a"]},
{"function": "rayleigh", "parameters": ["scale"]},
{"function": "triangular", "parameters": ["left", "mode", "right"]},
{"function": "uniform", "parameters": ["low", "high"]},
{"function": "vonmises", "parameters": ["mu", "kappa"]},
{"function": "wald", "parameters": ["mean", "scale"]},
{"function": "weibull", "parameters": ["a"]},
{"function": "zipf", "parameters": ["a"]}
}
Spectrum description configuration file¶
Spectrum description configuration file (spectrum_description.json) contains descriptions for standard solution and solid-state NMR experiments.
- List all available experiments:
In [51]:
nmrstarlib.nmrstarlib.list_spectrums()
CANCO
CANCOCX
CBCANH
CBCAcoNH
CCcoNH
HBHAcoNH
HNCA
HNCACB
HNCO
HNcaCO
HNcoCA
HNcoCACB
HSQC
HccoNH
NCA
NCACX
NCO
NCOCX
- List all available spectrum descriptions:
In [52]:
nmrstarlib.nmrstarlib.list_spectrum_descriptions()
{'CANCO': {'Labels': ['CA', 'N', 'CO-1'],
'MinNumberPeaksPerSpinSystem': 1,
'PeakDescriptions': [{'dimensions': ['CA', 'N', 'CO-1'], 'fraction': 1}]},
'CANCOCX': {'Labels': ['CA', 'N', 'CO-1', 'CX-1'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['CA', 'N', 'CO-1', 'CO-1'], 'fraction': 1},
{'dimensions': ['CA', 'N', 'CO-1', 'CA-1'], 'fraction': 1},
{'dimensions': ['CA', 'N', 'CO-1', 'CB-1'], 'fraction': 1},
{'dimensions': ['CA', 'N', 'CO-1', 'CG-1'], 'fraction': 1},
{'dimensions': ['CA', 'N', 'CO-1', 'CD-1'], 'fraction': 1},
{'dimensions': ['CA', 'N', 'CO-1', 'CE-1'], 'fraction': 1},
{'dimensions': ['CA', 'N', 'CO-1', 'CZ-1'], 'fraction': 1}]},
'CBCANH': {'Labels': ['CA/CB', 'H', 'N'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['CA', 'H', 'N'], 'fraction': 1},
{'dimensions': ['CB', 'H', 'N'], 'fraction': 0.95},
{'dimensions': ['CA', 'H+1', 'N+1'], 'fraction': 1},
{'dimensions': ['CB', 'H+1', 'N+1'], 'fraction': 0.95}]},
'CBCAcoNH': {'Labels': ['CA/CB', 'H+1', 'N+1'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['CA', 'H+1', 'N+1'], 'fraction': 1},
{'dimensions': ['CB', 'H+1', 'N+1'], 'fraction': 0.95}]},
'CCcoNH': {'Labels': ['CX-1', 'N', 'H'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['CA-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['CB-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['CG-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['CD-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['CE-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['CZ-1', 'N', 'H'], 'fraction': 1}],
'ResonanceLimit': {'ALA': ['H', 'N', 'CA', 'CB'],
'ARG': ['H', 'N', 'CA', 'CB', 'CG', 'CD', 'CZ'],
'ASN': ['H', 'N', 'CA', 'CB', 'CG'],
'ASP': ['H', 'N', 'CA', 'CB', 'CG'],
'CYS': ['H', 'N', 'CA', 'CB'],
'GLN': ['H', 'N', 'CA', 'CB', 'CG', 'CD'],
'GLU': ['H', 'N', 'CA', 'CB', 'CG', 'CD'],
'GLY': ['H', 'N', 'CA'],
'HIS': ['H', 'N', 'CA', 'CB'],
'ILE': ['H', 'N', 'CA', 'CB', 'CG1', 'CG2', 'CD1'],
'LEU': ['H', 'N', 'CA', 'CB', 'CG', 'CD1', 'CD2'],
'LYS': ['H', 'N', 'CA', 'CB', 'CG', 'CD', 'CE'],
'MET': ['H', 'N', 'CA', 'CB', 'CG', 'CE'],
'PHE': ['H', 'N', 'CA', 'CB'],
'SER': ['H', 'N', 'CA', 'CB'],
'THR': ['H', 'N', 'CA', 'CB', 'CG2'],
'TRP': ['H', 'N', 'CA', 'CB'],
'TYR': ['H', 'N', 'CA', 'CB'],
'VAL': ['H', 'N', 'CA', 'CB', 'CG1', 'CG2']}},
'HBHAcoNH': {'Labels': ['HA/HB-1', 'N', 'H'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['HA-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['HB-1', 'N', 'H'], 'fraction': 1}]},
'HNCA': {'Labels': ['H', 'N', 'CA'],
'MinNumberPeaksPerSpinSystem': 1,
'PeakDescriptions': [{'dimensions': ['H', 'N', 'CA'], 'fraction': 1},
{'dimensions': ['H', 'N', 'CA-1'], 'fraction': 1}]},
'HNCACB': {'Labels': ['H', 'N', 'CA/CB'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['H', 'N', 'CA'], 'fraction': 1},
{'dimensions': ['H', 'N', 'CB'], 'fraction': 0.95},
{'dimensions': ['H', 'N', 'CA-1'], 'fraction': 1},
{'dimensions': ['H', 'N', 'CB-1'], 'fraction': 0.95}]},
'HNCO': {'Labels': ['H', 'N', 'CO-1'],
'MinNumberPeaksPerSpinSystem': 1,
'PeakDescriptions': [{'dimensions': ['H', 'N', 'CO-1'], 'fraction': 1}]},
'HNcaCO': {'Labels': ['H', 'N', 'CO'],
'MinNumberPeaksPerSpinSystem': 1,
'PeakDescriptions': [{'dimensions': ['H', 'N', 'CO'], 'fraction': 1},
{'dimensions': ['H', 'N', 'CO-1'], 'fraction': 1}]},
'HNcoCA': {'Labels': ['H', 'N', 'CA'],
'MinNumberPeaksPerSpinSystem': 1,
'PeakDescriptions': [{'dimensions': ['H', 'N', 'CA-1'], 'fraction': 1}]},
'HNcoCACB': {'Labels': ['H', 'N', 'CA/CB-1'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['H', 'N', 'CA-1'], 'fraction': 1},
{'dimensions': ['H', 'N', 'CB-1'], 'fraction': 0.95}]},
'HSQC': {'Labels': ['H', 'N'],
'MinNumberPeaksPerSpinSystem': 1,
'PeakDescriptions': [{'dimensions': ['H', 'N'], 'fraction': 1}]},
'HccoNH': {'Labels': ['HX-1', 'N', 'H'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['HA-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['HB-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['HG-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['HD-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['HE-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['HH-1', 'N', 'H'], 'fraction': 1},
{'dimensions': ['HZ-1', 'N', 'H'], 'fraction': 1}],
'ResonanceLimit': {'ALA': ['H', 'N', 'HA', 'HB'],
'ARG': ['H', 'N', 'HA', 'HB2', 'HB3', 'HG2', 'HG3', 'HD2', 'HD3'],
'ASN': ['H', 'N', 'HA', 'HB2', 'HB3'],
'ASP': ['H', 'N', 'HA', 'HB2', 'HB3'],
'CYS': ['H', 'N', 'HA', 'HB2', 'HB3'],
'GLN': ['H', 'N', 'HA', 'HB2', 'HB3', 'HG2', 'HG3'],
'GLU': ['H', 'N', 'HA', 'HB2', 'HB3', 'HG2', 'HG3'],
'GLY': ['H', 'N', 'HA2', 'HA3'],
'HIS': ['H', 'N', 'HA', 'HB2', 'HB3'],
'ILE': ['H', 'N', 'HA', 'HB', 'HG12', 'HG13', 'HG2', 'HD1'],
'LEU': ['H', 'N', 'HA', 'HB2', 'HB3', 'HD1', 'HD2'],
'LYS': ['H', 'N', 'HA', 'HB2', 'HB3', 'HG2', 'HG3', 'HD2', 'HD3', 'HE2', 'HE3'],
'MET': ['H', 'N', 'HA', 'HB2', 'HB3', 'HG2', 'HG3', 'HE'],
'PHE': ['H', 'N', 'HA', 'HB2', 'HB3'],
'SER': ['H', 'N', 'HA', 'HB2', 'HB3'],
'THR': ['H', 'N', 'HA', 'HB', 'HG2'],
'TRP': ['H', 'N', 'HA', 'HB2', 'HB3'],
'TYR': ['H', 'N', 'HA', 'HB2', 'HB3'],
'VAL': ['H', 'N', 'HA', 'HB', 'HG1', 'HG2']}},
'NCA': {'Labels': ['N', 'CA'],
'MinNumberPeaksPerSpinSystem': 1,
'PeakDescriptions': [{'dimensions': ['N', 'CA'], 'fraction': 1}]},
'NCACX': {'Labels': ['N', 'CA', 'CX'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['N', 'CA', 'CO'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CA'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CB'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CG'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CD'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CE'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CZ'], 'fraction': 1}]},
'NCO': {'Labels': ['N', 'CO-1'],
'MinNumberPeaksPerSpinSystem': 1,
'PeakDescriptions': [{'dimensions': ['N', 'CO-1'], 'fraction': 1}]},
'NCOCX': {'Labels': ['N', 'CO-1', 'CX-1'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['N', 'CO-1', 'CA-1'], 'fraction': 1},
{'dimensions': ['N', 'CO-1', 'CB-1'], 'fraction': 1},
{'dimensions': ['N', 'CO-1', 'CG-1'], 'fraction': 1},
{'dimensions': ['N', 'CO-1', 'CD-1'], 'fraction': 1},
{'dimensions': ['N', 'CO-1', 'CE-1'], 'fraction': 1},
{'dimensions': ['N', 'CO-1', 'CZ-1'], 'fraction': 1}]}}
- List specific spectrum descriptions:
In [53]:
nmrstarlib.nmrstarlib.list_spectrum_descriptions("HNcoCACB", "NCACX")
{'HNcoCACB': {'Labels': ['H', 'N', 'CA/CB-1'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['H', 'N', 'CA-1'], 'fraction': 1},
{'dimensions': ['H', 'N', 'CB-1'], 'fraction': 0.95}]}}
{'NCACX': {'Labels': ['N', 'CA', 'CX'],
'MinNumberPeaksPerSpinSystem': 2,
'PeakDescriptions': [{'dimensions': ['N', 'CA', 'CO'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CA'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CB'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CG'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CD'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CE'], 'fraction': 1},
{'dimensions': ['N', 'CA', 'CZ'], 'fraction': 1}]}}
Adding a custom experiment description and simulating peak list based on it. Custom spectrum description can be added in several ways:
- Create additional json configuration with spectrum description and updating
SPECTRUM_DESCRIPTIONS
dict. Content of custom_spectrum_description.json.
{
"NCACX_custom": {
"Labels": ["N", "CA", "CX"],
"MinNumberPeaksPerSpinSystem": 2,
"PeakDescriptions": [
{"fraction": 1, "dimensions": ["N", "CA", "CO"]},
{"fraction": 1, "dimensions": ["N", "CA", "CA"]},
{"fraction": 1, "dimensions": ["N", "CA", "CB"]}
]
}
}
In [54]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToPeakList
from nmrstarlib.noise import NoiseGenerator
# update SPECTRUM_DESCRIPTIONS
nmrstarlib.nmrstarlib.update_constants(spectrum_descriptions_cfg="path/to/custom_spectrum_description.json")
# create parameters dictionary for random normal distribution
parameters = {"H_loc": [None, None], "C_loc": [0, 0], "N_loc": [0, 0],
"H_scale": [None, None], "C_scale": [0.01, 0.05], "N_scale": [0.01, 0.05]}
# create random normal noise generator
random_normal_noise_generator = NoiseGenerator(parameters)
converter = Converter(StarFileToPeakList(from_path="18569", to_path="out/18569_NCACX_custom.txt",
from_format="nmrstar", to_format="sparky",
spectrum_name="NCACX_custom",
plsplit=(70,30),
noise_generator=random_normal_noise_generator))
converter.convert()
- Define dictionary with new spectrum description and update SPECTRUM_DESCRIPTIONS
dict.
In [55]:
from nmrstarlib.converter import Converter
from nmrstarlib.translator import StarFileToPeakList
from nmrstarlib.noise import NoiseGenerator
custom_experiment_type = {
"NCACX_custom": {
"Labels": ["N", "CA", "CX"],
"MinNumberPeaksPerSpinSystem": 2,
"PeakDescriptions": [
{"fraction": 1, "dimensions": ["N", "CA", "CO"]},
{"fraction": 1, "dimensions": ["N", "CA", "CA"]},
{"fraction": 1, "dimensions": ["N", "CA", "CB"]}
]
}
}
# update SPECTRUM_DESCRIPTION
nmrstarlib.nmrstarlib.SPECTRUM_DESCRIPTIONS.update(custom_experiment_type)
# create parameters dictionary for random normal distribution
parameters = {"H_loc": [0, 0], "C_loc": [None, None], "N_loc": [0, 0],
"H_scale": [0.001, 0.005], "C_scale": [None, None], "N_scale": [0.01, 0.05]}
# create random normal noise generator
random_normal_noise_generator = NoiseGenerator(parameters)
# Using valid BMRB id to access file from URL: from_path="18569"
converter = Converter(StarFileToPeakList(from_path="18569", to_path="out/18569_NCACX_custom.txt",
from_format="nmrstar", to_format="sparky",
spectrum_name="NCACX_custom",
plsplit=(70,30),
noise_generator=random_normal_noise_generator))
converter.convert()
Visualizing chemical shifts values¶
Chemical shifts values can be visualized using the nmrstarlib.csviewer
Chemical Shifts Viewer module.
- Visualize all available chemical shifts for all amino acids.
In [56]:
from nmrstarlib.csviewer import CSViewer
csviewer = CSViewer(from_path="18569", filename="out/18569_chem_shifts_all", csview_format="png")
csviewer.csview(view=False)
nmrstarlib.csviewer output example:

- Visualize CA, CB, CG, and CG2 chemical shifts for specific amino acids.
In [57]:
from nmrstarlib.csviewer import CSViewer
csviewer = CSViewer(from_path="18569", amino_acids=["GLU", "THR"], atoms=["CA", "CB", "CG", "CG2"],
filename="out/18569_chem_shifts_SER_THR_CA_CB_CG_CG2", csview_format="png")
csviewer.csview(view=False)
nmrstarlib.csviewer output example:
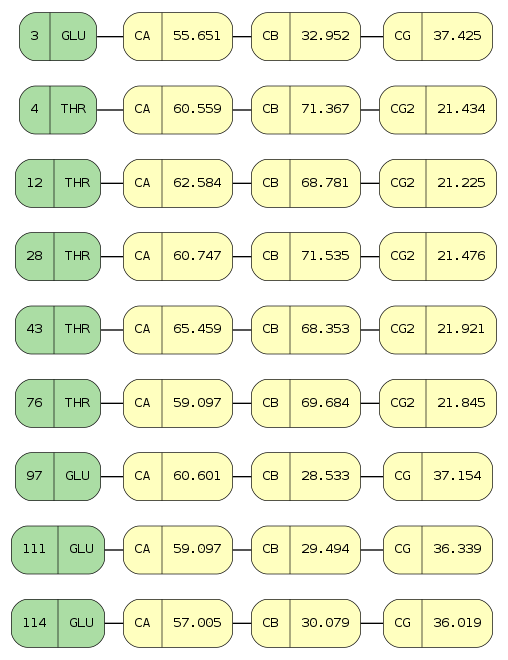

- Visualize specific atoms for specific amino acids.
In [58]:
from nmrstarlib.csviewer import CSViewer
csviewer = CSViewer(from_path="18569", amino_acids_and_atoms={"GLU": ["CA", "CB"], "THR": ["HA", "HB"]},
filename="out/18569_chem_shifts_GLU_CA_CB_THR_HA_HB", csview_format="png")
csviewer.csview(view=False)
nmrstarlib.csviewer output example:
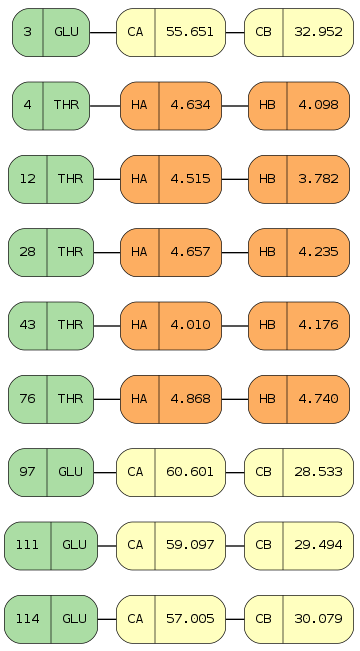
Command Line Interface¶
- Command Line Interface functionality:
- Convert from the NMR-STAR file format into its equivalent JSON file format and vice versa.
- Create simulated peak list files using chemical shift and assignment information.
- Visualize assigned chemical shift values.
In [59]:
! python3 -m nmrstarlib --help
nmrstarlib command-line interface
Usage:
nmrstarlib -h | --help
nmrstarlib --version
nmrstarlib convert (<from-path> <to-path>) [--from-format=<format>] [--to-format=<format>] [--bmrb-url=<url> | --pdb-url=<url>] [--nmrstar-version=<version>] [--verbose]
nmrstarlib csview <starfile-path> [--aa=<aa>] [--at=<at>] [--aa-at=<aa-at>] [--csview-outfile=<path>] [--csview-format=<format>] [--bmrb-url=<url> | --pdb-url=<url>] [--nmrstar-version=<version>] [--verbose] [--show]
nmrstarlib plsimulate (<from-path> <to-path> <spectrum>) [--from-format=<format>] [--to-format=<format>] [--plsplit=<%>] [--distribution=<func>] [--seed=<value>] [--H=<value>] [--C=<value>] [--N=<value>] [--bmrb-url=<url> | --pdb-url=<url>] [--nmrstar-version=<version>] [--spectrum-descriptions=<path>] [--verbose]
Options:
-h, --help Show this screen.
--version Show version.
--verbose Print what files are processing.
--show Display chemical shifts image generated by 'csview' command by default image viewer.
--from-format=<format> Input file format, available formats: nmrstar, json [default: nmrstar].
--to-format=<format> Output file format, available formats: nmrstar, json [default: json].
--nmrstar-version=<version> Version of NMR-STAR format to use, available: 2, 3 [default: 3].
--bmrb-url=<url> URL to BMRB interface [default: http://rest.bmrb.wisc.edu/bmrb/NMR-STAR3/].
--pdb-url=<url> URL to PDB interface [default: https://files.rcsb.org/view/].
--aa=<aa> Comma-separated amino acid three-letter codes (e.g. --aa=ALA,SER).
--at=<at> Comma-separated BMRB atom codes (e.g. --at=CA,CB).
--aa-at=<aa-at> Amino acid three-letter codes (keys) and corresponding atoms (values) (e.g. --aa-at=ALA-CA,CB:LYS-CB,CG,CD).
--csview-outfile=<path> Where to save chemical shifts table.
--csview-format=<format> Format to which save chemical shift table [default: svg].
--plsplit=<%> How to split peak list into chunks by percent [default: 100].
--spectrum-descriptions=<path> Path to custom spectrum descriptions file.
--distribution=<func> Statistical distribution function [default: normal].
--seed=<value> Integer value used to initialize a pseudorandom number generator during peak list simulation.
--H=<value> Statistical distribution parameter(s) for H dimension.
--C=<value> Statistical distribution parameter(s) for C dimension.
--N=<value> Statistical distribution parameter(s) for N dimension.
CLI Converting NMR-STAR files in bulk¶
CLI one-to-one file conversions¶
- Convert from a local file in NMR-STAR format to a local file in JSON format:
In [60]:
! python3 -m nmrstarlib convert bmr18569.str out/bmr18569.json \
--from-format=nmrstar --to-format=json
- Convert from a local file in JSON format to a local file in NMR-STAR format:
In [61]:
! python3 -m nmrstarlib convert bmr18569.json out/bmr18569.str \
--from-format=json --to-format=nmrstar
- Convert from a compressed local file in NMR-STAR format to a compressed local file in JSON format:
In [62]:
! python3 -m nmrstarlib convert bmr18569.str.gz out/bmr18569.json.gz \
--from-format=nmrstar --to-format=json
- Convert from a compressed local file in JSON format to a compressed local file in NMR-STAR format:
In [63]:
! python3 -m nmrstarlib convert bmr18569.json.gz out/bmr18569.str.gz \
--from-format=json --to-format=nmrstar
- Convert from a uncompressed URL file in NMR-STAR format to a compressed local file in JSON format:
In [64]:
! python3 -m nmrstarlib convert 18569 out/bmr18569.json.bz2 \
--from-format=nmrstar --to-format=json
Note
See nmrstarlib.converter for full list of available
conversions.
CLI many-to-many files conversions¶
- Convert from a directory of files in NMR-STAR format to a directory of files in JSON format:
In [65]:
! python3 -m nmrstarlib convert starfiles_dir_nmrstar out/starfiles_dir_json \
--from-format=nmrstar --to-format=json
- Convert from a directory of files in JSON format to a directory of files in NMR-STAR format:
In [66]:
! python3 -m nmrstarlib convert starfiles_dir_json out/starfiles_dir_nmrstar \
--from-format=json --to-format=nmrstar
- Convert from a directory of files in NMR-STAR format to a zip archive of files in JSON format:
In [67]:
! python3 -m nmrstarlib convert starfiles_dir_nmrstar out/starfiles_json.zip \
--from-format=nmrstar --to-format=json
- Convert from a compressed tar archive of files in JSON format to a directory of files in NMR-STAR format:
In [68]:
! python3 -m nmrstarlib convert starfiles_json.tar.gz out/starfiles_dir_nmrstar \
--from-format=json --to-format=nmrstar
- Convert from a zip archive of files in NMR-STAR format to a compressed tar archive of files in JSON format:
In [69]:
! python3 -m nmrstarlib convert starfiles_nmrstar.zip out/starfiles_json.tar.bz2 \
--from-format=nmrstar --to-format=json
Note
See nmrstarlib.converter for full list of available
conversions.
CLI Creating simulated peak list files from NMR-STAR files in bulk¶
CLI one-to-one file simulations¶
- Creating a zero-variance HNcoCACB peak list file in sparky-like format from local NMR-STAR formatted file (bmr18569.str):
In [70]:
! python3 -m nmrstarlib plsimulate bmr18569.str out/18569_HNcoCACB.txt HNcoCACB \
--from-format=nmrstar --to-format=sparky
- Creating a HNcoCACB peak list file in sparky-like format and adding noise values to peak dimensions from a single source of variance, i.e. 100% of peaks will have chemical shift values adjusted using noise values from the defined random normal distribution (note that we can use 18569 BMRB id instead of local file):
In [71]:
! python3 -m nmrstarlib plsimulate 18569 out/18569_HNcoCACB_ssv_HCN.txt HNcoCACB \
--from-format=nmrstar --to-format=sparky \
--H=0,0.001 --N=0,0.01 --C=0,0.01
- Creating a HNcoCACB peak list file in sparky-like format and adding noise values to peak dimensions from a single source of variance, i.e. 100% of peaks will have chemical shift values adjusted using noise values from the defined chisquare distribution for degrees of freedom equal to 5:
In [72]:
! python3 -m nmrstarlib plsimulate 18569 out/18569_HNcoCACB_ssv_HCN_chi2.txt HNcoCACB \
--from-format=nmrstar --to-format=sparky \
--H=5 --N=5 --C=5 --distribution=chisquare
- Creating a HNcoCACB peak list file in sparky-like format and adding noise values to H and N peak dimensions but not C peak dimension from a single source of variance, i.e. 100% of peaks will have chemical shift values adjusted using noise values from the defined random normal distribution (note that we can use compressed bmr18569.str.gz file):
In [73]:
! python3 -m nmrstarlib plsimulate bmr18569.str.gz out/18569_HNcoCACB_ssv_HN.txt HNcoCACB \
--from-format=nmrstar --to-format=sparky \
--H=0,0.001 --N=0,0.01
- Creating a HNcoCACB peak list file in sparky-like format and adding
noise values to peak dimensions from two sources of variance, i.e.
chemical shift values will be adjusted using noise values
from two random normal distributions. In order to specify two sources
of variance, we need to provide how we want to split our peak list and
provide statistical distribution parameters for both distributions. Let’s
say we want 70 % of peaks to have a smaller variance in H and N dimensions
and 30 % of peaks to have a larger variance in H and N dimensions. Note
that values per split are separated by
:and then parameters are separated by,.
In [74]:
! python3 -m nmrstarlib plsimulate 18569 out/18569_HNcoCACB_tsv_HN.txt HNcoCACB \
--from-format=nmrstar --to-format=sparky \
--plsplit=70,30 --H=0:0,0.001:0.005 --N=0:0,0.01:0.05
Note
See nmrstarlib.converter for full list of available one-to-one and many-to-many
input and output formats.
CLI many-to-many files simulations¶
- Simulate zero-variance HNcoCACB peak lists from a directory of NMR-STAR formatted files to a directory of peak list files:
In [75]:
! python3 -m nmrstarlib plsimulate starfiles_dir_nmrstar out/peaklists_dir HNcoCACB \
--from-format=nmrstar --to-format=sparky
- Simulate HNcoCACB peak lists from a directory of NMR-STAR formatted files to a zip archive of peak list files, add random normal noise values to H and N peak dimensions:
In [76]:
! python3 -m nmrstarlib plsimulate starfiles_dir_nmrstar out/peaklists.zip HNcoCACB \
--from-format=nmrstar --to-format=sparky --H=0,0.001 --N=0,0.01
- Simulate NCACX peak lists from a directory of NMR-STAR formatted files to a tar.gz archive of peak list files, add random normal noise values to C and N peak dimensions using two sources of variance, 70 % of peaks will have smaller variance, 30 % of peaks will have larger variance:
In [77]:
! python3 -m nmrstarlib plsimulate starfiles_dir_nmrstar out/peaklists.tar.gz NCACX \
--from-format=nmrstar --to-format=sparky --plsplit=70,30 \
--C=0:0,0.01:0.05 --N=0:0,0.01:0.07
Note
See nmrstarlib.converter for full list of available one-to-one and many-to-many
input and output formats.
CLI Visualizing chemical shift values¶
- Visualize chemical shift values for the entire sequence:
In [78]:
! python3 -m nmrstarlib csview 18569 --csview-outfile=out/18569_chem_shifts_all \
--csview-format=png

- Visualize CA, CB, CG, and CG2 chemical shift values for GLU and THR amino acid residues:
In [79]:
! python3 -m nmrstarlib csview 18569 --aa=GLU,THR --at=CA,CB,CG,CG2 \
--csview-outfile=out/18569_chem_shifts_GLU_THR_CA_CB_CG_CG2 \
--csview-format=png

- Visualize specific atoms for specific amino acids.
In [80]:
! python3 -m nmrstarlib csview 18569 --aa-at=GLU-CA,CB:THR-HA,HB \
--csview-outfile=out/18569_chem_shifts_GLU_CA_CB_THR_HA_HB \
--csview-format=png
nmrstarlib.csviewer output example:
